Research Vision
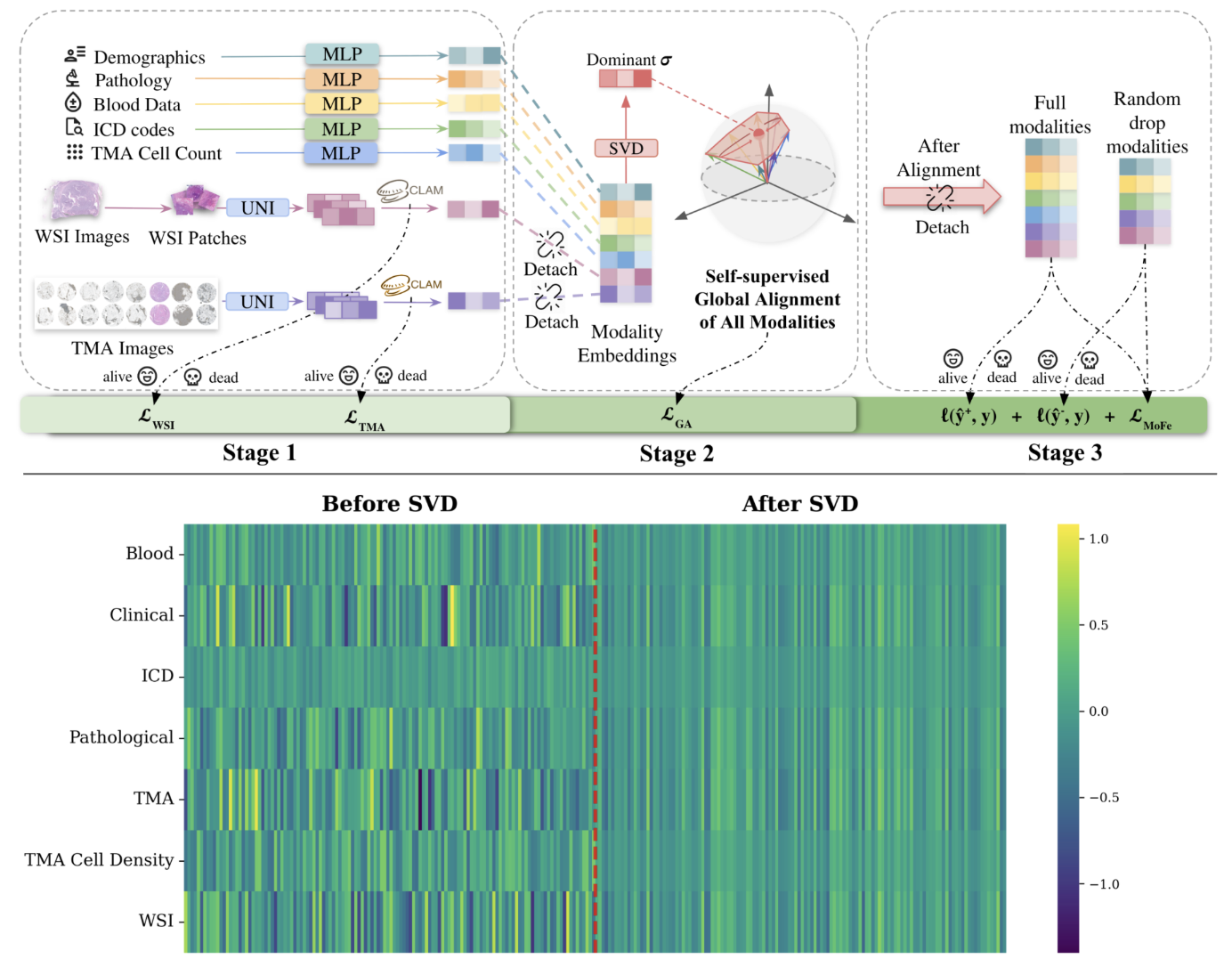
I study learning dynamics in multimodal biomedical AI systems, with a focus on survival prediction and risk modeling under heterogeneous and missing-modality settings. Using data such as histopathology images, structured clinical variables, and molecular signals, I investigate how architectural design and optimization strategies transform partially observed, noisy inputs into stable and learnable decision representations.
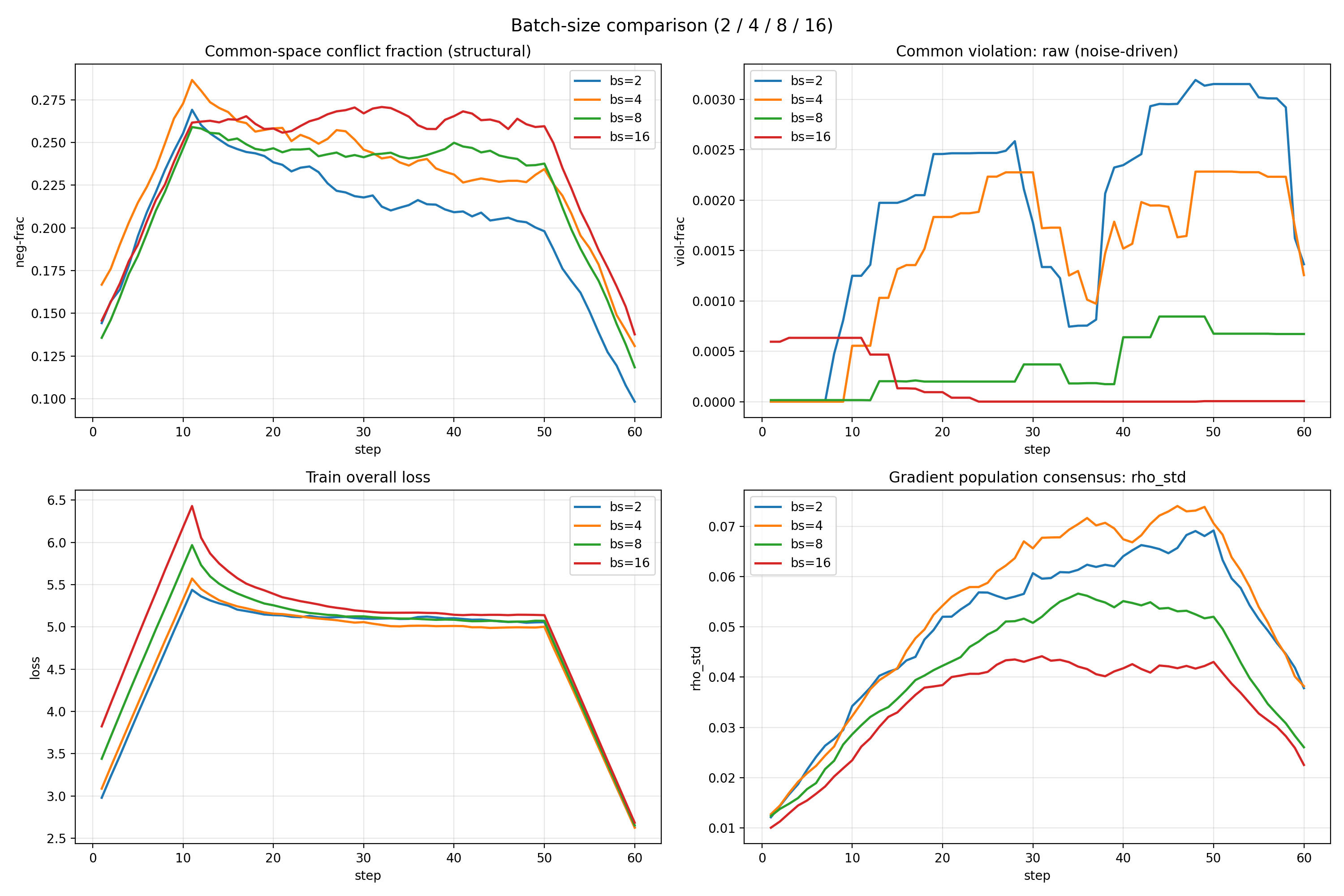
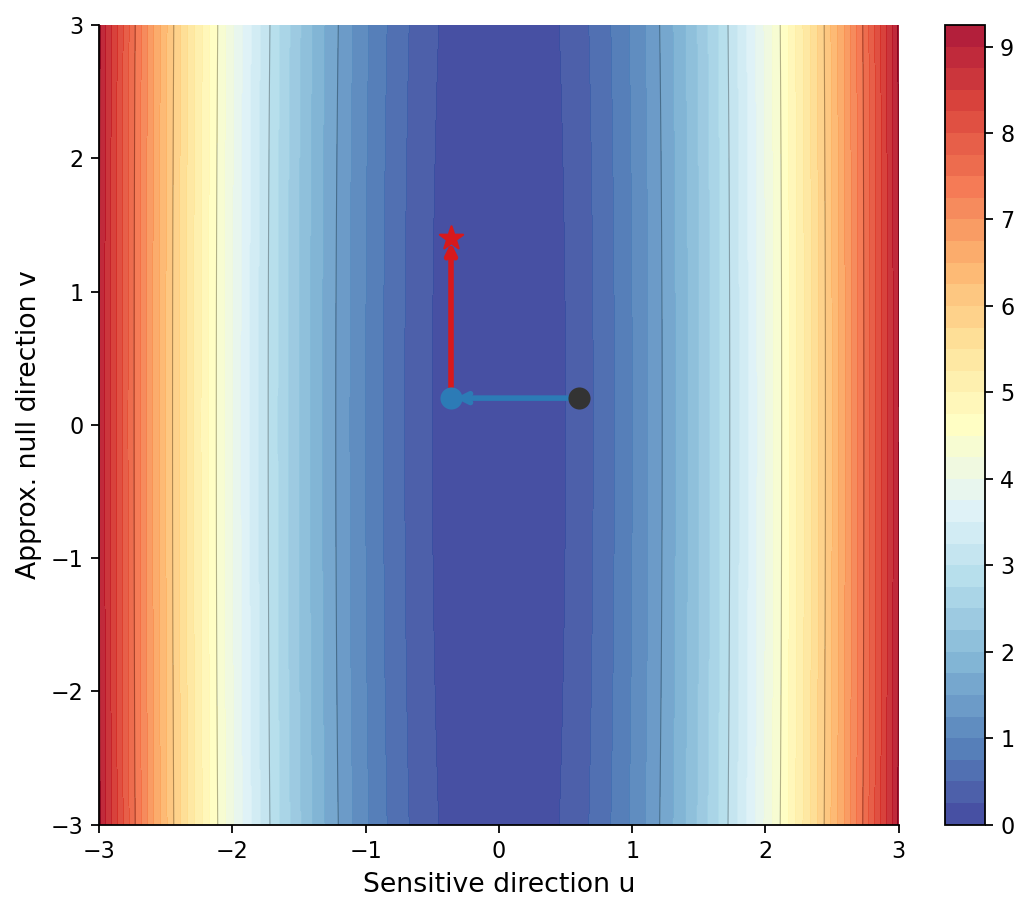
My research spans both molecular and imaging scales. At the molecular level, I fine-tune protein foundation models using more than two million sequences and inject post-translational modification (PTM) information to enhance structural and functional representation learning. At the imaging level, I analyze gradient interactions across multi-task pathology heads, characterizing structural subspace conflicts, stochastic batch-induced noise conflicts, and dominance imbalance among tasks. By studying how different gradient strategies reshape shared backbones, I seek optimization mechanisms that preserve task-relevant geometry while mitigating destructive interference.
Rather than viewing multimodal learning as simple fusion, I see machine learning more broadly as a mathematical process that organizes complex, noisy real-world information into structured representations capable of supporting reliable decisions. My goal is to understand how model structure and optimization dynamics shape this transformation—how they filter noise, integrate heterogeneous signals, and amplify task-relevant information within high-dimensional spaces.
Biomedical systems, with their multi-scale interactions, partial observability, and inherent uncertainty, make these structural challenges explicit. By studying how useful signals emerge and stabilize under such conditions, I aim to design AI systems that more deliberately and effectively extract what truly matters from data.